Quantum Chemistry and Protein structures
Thought for 27s
Title: AI and Quantum Chemistry Just Made Protein Structures Sharper
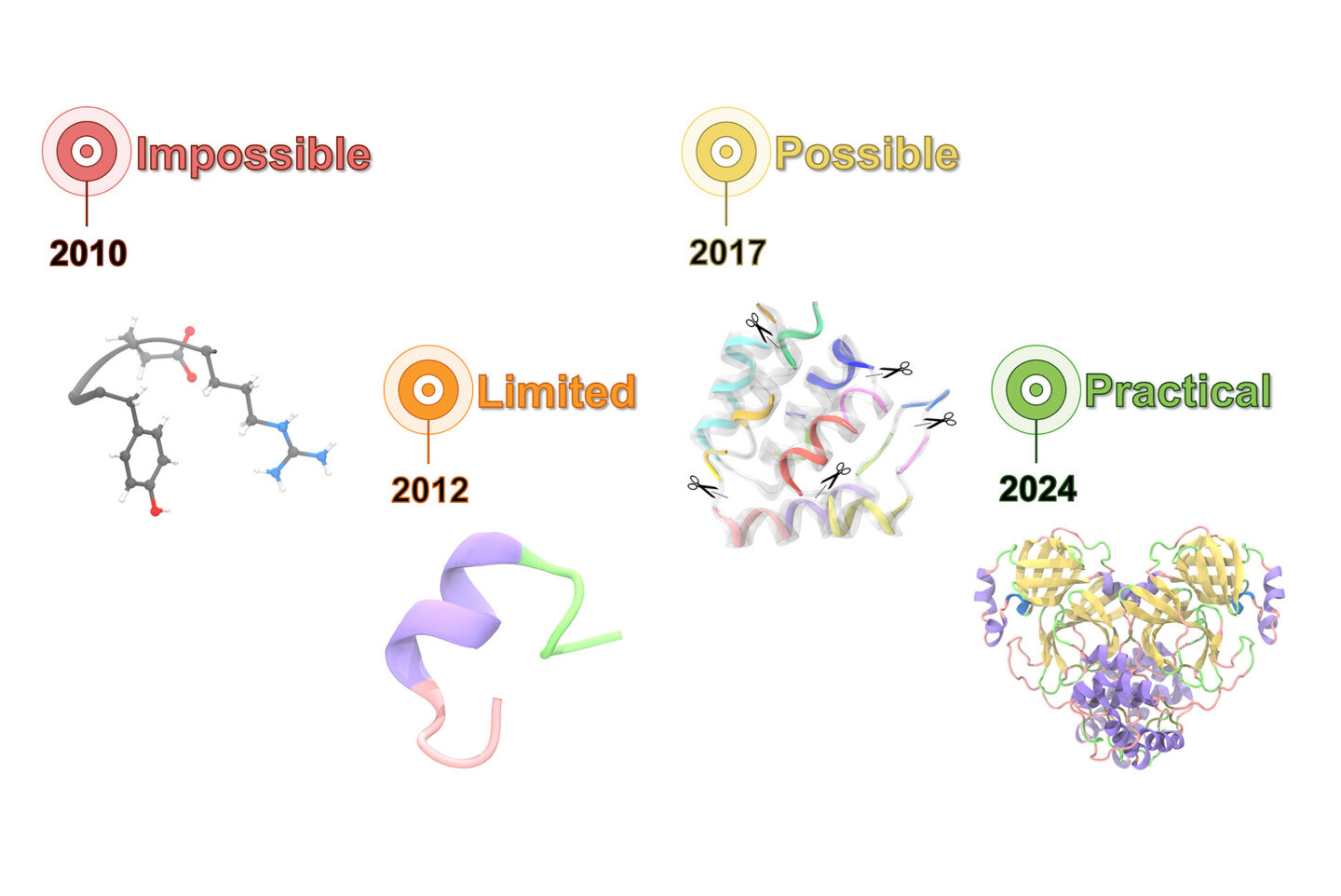
A lot of people think the protein structure revolution is mostly finished because we can predict folds quickly. In the last couple of weeks, a different kind of advance made the rounds, and it matters for the parts of biology where small details change the story. Berkeley Lab reported a new tool called AQuaRef that refines experimentally measured protein structures using a mix of quantum mechanical ideas and modern AI, aiming to place atoms and even protons more accurately than standard refinement methods.
Here is the problem it targets. Experimental techniques like cryo EM and X ray crystallography give you strong evidence about where atoms should be, but turning that evidence into a final atomic model depends on chemical “restraints” that come from libraries of known structures and known chemistry. Those libraries do not fully capture the subtle noncovalent interactions that actually hold proteins together, and they struggle when the chemistry is unusual or the data is lower resolution. In principle, quantum mechanical calculations can provide that missing realism, but they are usually too expensive for large biomolecules.
AQuaRef tries to make that quantum level guidance practical by using a machine learned interatomic potential that mimics quantum behavior at a much lower computational cost. In the Nature Communications paper, the authors report that across dozens of cryo EM and X ray structures, their approach improved geometric quality while keeping an equal or better fit to the experimental data. They also highlight a concrete win that many structural biologists care about: better proton placement in a notoriously tricky human protein called DJ 1, linked to some forms of parkinsonism.
The other reason this is a meaningful news story is that it plugs into workflows people already use. Berkeley Lab notes that AQuaRef is part of Phenix, a widely used software suite for macromolecular structure determination. That means this is not only a research demo. It is positioned as an upgrade to the refinement step many labs run every day when they solve new structures.
If this direction holds, it changes what “solving a structure” can mean. You can have a model that looks correct at the ribbon level, yet misses the local chemistry that controls binding, catalysis, and stability. Tools that sharpen those details are exactly what you want for drug design and for understanding disease mechanisms that depend on small, local interactions rather than large, global shapes.
Sources
https://newscenter.lbl.gov/2026/03/10/cracking-the-code-using-ai-to-solve-difficult-to-map-proteins/
https://www.nature.com/articles/s41467-025-64313-1
https://phenix-online.org/
- Get link
- X
- Other Apps
Comments
Post a Comment